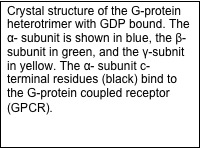
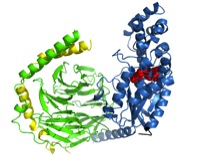
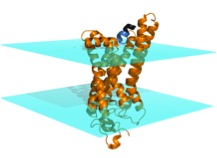
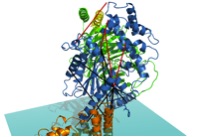
Rhodopsin is a G-protein-coupled receptor (GPCR), which are integral membrane proteins with seven transmembrane helices. When activated by an extracellular affecter, GPCRs bind to G-proteins, catalyzing the release of GDP from the G-protein. The alpha subunit of the heterotrimeric G-protein (Galpha) can then interact with intracellular partners and propagate the signal into the cell. The protein arrestin binds to GPCRs in order to regulate their activity.
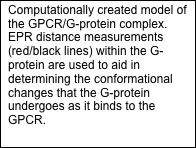
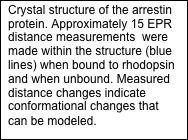
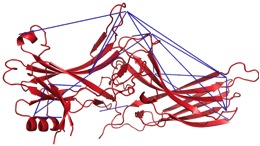
Crystal structures exist for the G-protein, as well as activated rhodopsin bound to the C-terminal tail of Galpha, but not for the rhodopsin/G-protein complex. In addition, crystal structures exist for arrestin proteins, but not in complex with a GPCR. Currently, we use Rosetta in conjunction with EPR distance and mobility measurements in order to model the rhodopsin/G-protein complex and conformational changes occurring as the G-protein binds to rhodopsin. EPR distance measurements are also used to model the conformational changes of arrestin as it binds to rhodopsin.




Alumni Project Members: Nathan Alexander
