2026 April 14: VUStruct is back online!
Over recent months, Vanderbilt's ACCRE cluster has been dramtically upgraded and the VUStruct web frontend had to be reworked.
If you have any difficulties using VUStruct, please Get in touch.Evolutionary Constraint: CONSURF and COSMIS
Residue positions which vary infrequently across species exhibit evolutionary conservation and this is already an input to the clinic via GERP(Cooper et al. 2005) and GERP++(Davydov et al. 2010) scores.
As structural biologists, we are interested not only in conservation for specific amino acids in a sequence, but also in the conservation, and visualization, of the surrounding 3D protein context. ConSurf(Ashkenazy et al. 2016) colors every residue based on rates of evolution from phylogenetic tree analysis (Pupko et al. 2002 ).
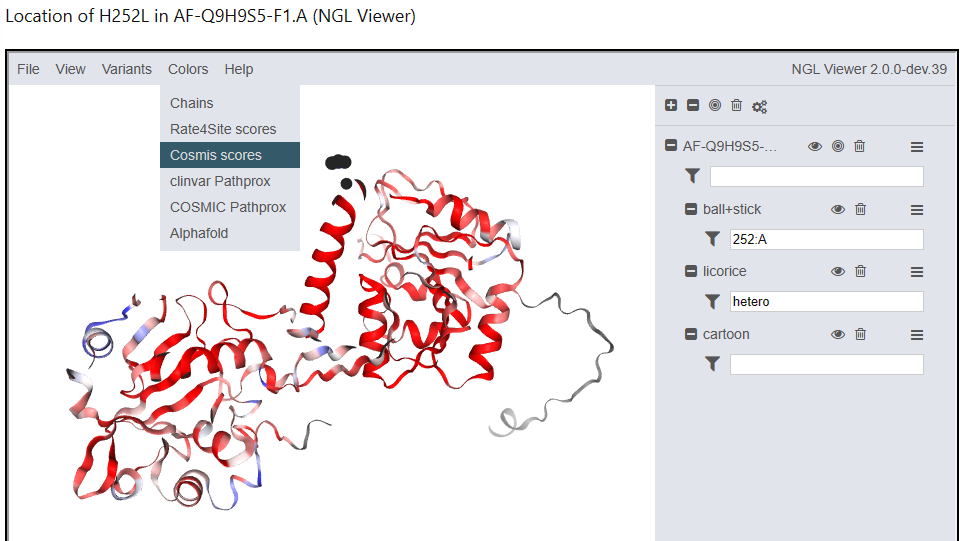
The Contact Set MISsense tolerance algorithm, COSMIS (Li et al. 2022) was developed by Bian Li of the Capra Lab. In contrast to Consurf, which quantifies evolutionary constraint based on analysis of sequence alignments across diverse species, COSMIS quantifies evolutionary constraint over more recent evolution by analyzing patterns of genetic variation across diverse human populations. It also leverages the 3D spatial context of proteins to better estimate the evolutionary constraint on protein positions of interest. The figure below is an example depiction of COSMIS scores.